
Features
Genetics
Research
Tunatech trials boast 100% success in early sturgeon sex assignment
The German biotechnology company touts a “game–changer” diagnostic technique that could enable extensive savings on costs of caviar production.
December 13, 2022 By Christopher Bridges, TunaTech GmbH
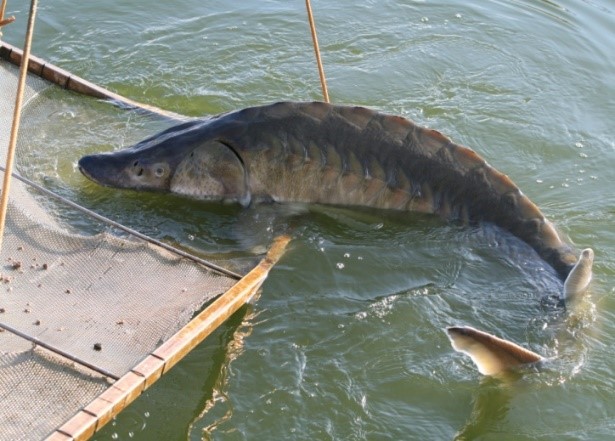 14-year-old adult sturgeon broodstock (Landesfischerei Anstalt) in Born, Germany (Photo: TunaTech Aquaculture)
14-year-old adult sturgeon broodstock (Landesfischerei Anstalt) in Born, Germany (Photo: TunaTech Aquaculture) Throughout the world many sturgeon hatcheries (of which there are more than 2,000) are faced with a problem that there are no externally visible sexual dimorphisms which enables one to distinguish between males and females. Fish must be kept for up to 10-14 years before sex can be determined using ultrasonic probes. Early sexual identification can significantly decrease the production costs by removing unwanted males from the production line and increasing the number of females held within the facility.
For us, the search for a sex specific molecular marker, the “Holy Grail,” for sturgeon propagation management has been long and began with numerous Bachelor and Master’s studies at the Heinrich Heine University in Düsseldorf, Germany. The early results were not encouraging and the use of random primer sets to try and distinguish between males and females at the molecular level proved to be ineffective.
Work continued with the Landesfischeri Anstalt in the city of Born who had a supply of mature sturgeon and were engaged in a restocking programme in the river Elbe. Their hatchery was generating large numbers of fingerlings for the various river systems. At this time, myself (Chris Bridges), Florian Borutta and Stephan Schulz began looking for international collaboration and funding.
Our first attempt was not too successful but after the founding of the spin-off company, TunaTech, from the University of Düsseldorf, together with CEO Shukry Na’amnieh, we were able to access national funds for our transcriptome analysis of the genome of adult male and female sturgeon samples. These were collected from the Aquatir sturgeon farm in Moldova with the help of aquaculture consultant, Dietmar Firzlaff (aquaFUTURE e.K.). These studies yielded an immense amount of data and then screening began together with the Informatics department at the University of Düsseldorf in the search for the veritable “needle in a haystack.” A number of likely marker combinations were found but these were not verifiable and failed proof of concept tests.
The major breakthrough came when another team of European scientists from STURGEoNOMICS, an EU-funded research project, published the results of their genome analysis work in 2020, indicating a female-specific genome region responsible for sex determination in the sturgeon. Using information from this paper, together with methodology implicit from our previous studies, it was possible to develop a commercially viable test. This involved identifying primer pairs which in qPCR analysis produced 100 per cent identification of female fish.
Through the range of samples available to us (i.e. blood, tissue and mucus samples of sexually mature Belugas (Huso huso)) Russian Sturgeons (Acipenser gueldenstaedtii) and Sterlets (Acipenser ruthenus) with known sex from Moldova and juveniles from the north of Germany we were able to start a rigorous proof of concept analysis.
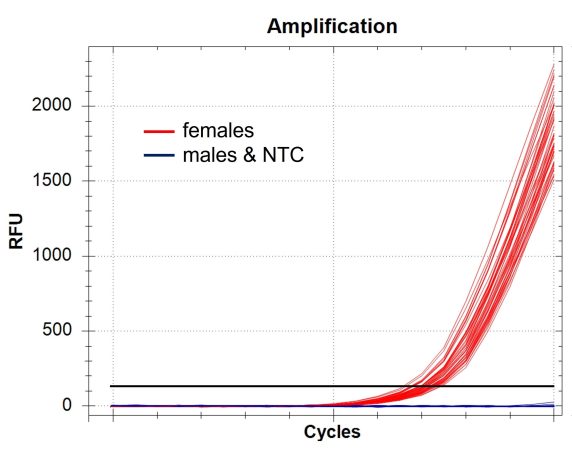
Exemplary presentation of the results of commercial DNA analysis using qPCR of fin clips and blood taken from Siberian Sturgeon, Russian Sturgeon and Beluga. The clear positive RFU signal is only found within the females. Figure displays 46 female and 46 male samples as well as 4 NTCs(Relative Fluorescence Units, RFU; Non Template Control, NTC). (Photo: TunaTech Aquaculture)
Thorough testing in the laboratory of samples from multiple sturgeon species ranging from adult fish of 15 years of age down to juvenile sturgeons of approximately four months have indicated that the marker can be detected even at very early stages in the development.
To simplify matters for possible on-site users, further research and development work went on o developed an easy colour test for sex discrimination which does not involve polymerase chain reaction (PCR) or quantitative polymerase chain reaction (qPCR) analysis but just simple incubation of extracted DNA as shown below and is based on the LAMP (Loop-Mediated Isothermal Amplification) technology. This alternative technology can be applied where rapid onsite results are required without extensive laboratory backup or logistics.
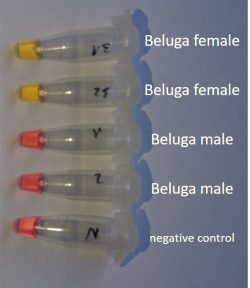
Colour test results of a LAMP determination for Beluga sturgeon (Photo: TunaTech Aquaculture)
The methodology applied is that individual hatcheries are required to obtain genomic tissue samples either as a fin-clip or using a tissue sampling unit (TSU) from each fish. To identify individual fish after testing then a marking procedure must be applied. The method of choice is a passive integrated transponder (PIT) tag which allows high throughput identification using external scanning. TSU’s and PIT tags are available from numerous commercial suppliers. Alternatives are available but not as efficient.
DNA is then extracted using a standard procedure – specifically adapted to fish fin material – a sex-determining gene fragment is then amplified by qPCR, and then the sexes are assigned to the respective samples in a software-supported and automatic manner.
Having established the correct protocol and methodologies to be used in the early sturgeon sex discrimination (ESSD) we set out to obtain field samples from hatcheries under true commercial conditions. In the initial phase the team has offered commercial producers the possibility of sending blind samples for testing from fish of known sex as proof of concept. The first seven specimens of Acipenser baeri, from an Italian farm (Emanuele Durigon in Quinto di Treviso, Italy, Dr. Vanni Durigon), which were seven years old and were sent for testing and using the protocol outlined above revealed four females and three males agreeing exactly with the predetermined sex.
In Germany the first pilot commercial trial of 16 fin clip samples came from a farm in Hessen (Fischzucht Rhönforelle GmbH & Co. KG). These fish consisted of Russian sturgeon (Acipenser gueldenstaedtii) broodstock which were 12-15 years of age.
The results indicated that five fish were females. Initially the farm confirmed the first four samples as females but thought the fifth was a male. However, after ESSD analysis, they checked this result by ultrasound and found that it was indeed a female. After this successful initial pilot study, a large-scale order from the same farm has now been processed and analyzed.
This game changer technology for sturgeon hatcheries throughout the world can make considerable savings possible, firstly eliminating the feeding of unwanted males and secondly increasing the stocking density with females. High throughput tagging and DNA sampling can be achieved for large numbers of fingerlings/immature fish for rapid separation of the required individuals.
Further information can be obtained at info@tunatech.de.
 Chris Bridges is co-founder of TUNATECH GmbH, researching diagnostics for fish reproduction in both marine and freshwater. Bridges is a marine biologist who studied in Liverpool, England before taking up a post at the Heinrich Heine University of Düsseldorf, Germany. He also has over 40 years of experience working in fish biology worldwide and the last 20 years on Atlantic bluefin tuna aquaculture.
Chris Bridges is co-founder of TUNATECH GmbH, researching diagnostics for fish reproduction in both marine and freshwater. Bridges is a marine biologist who studied in Liverpool, England before taking up a post at the Heinrich Heine University of Düsseldorf, Germany. He also has over 40 years of experience working in fish biology worldwide and the last 20 years on Atlantic bluefin tuna aquaculture.
Print this page
Advertisement
- Hatchery stocking of black tiger shrimp to help revive Philippine industry
- Virginia Tech opens new seafood research centre





