
Features
Breeding
Disease Management
Research
Clam genome project targets QPX disease
May 1, 2020 By Lynn Fantom
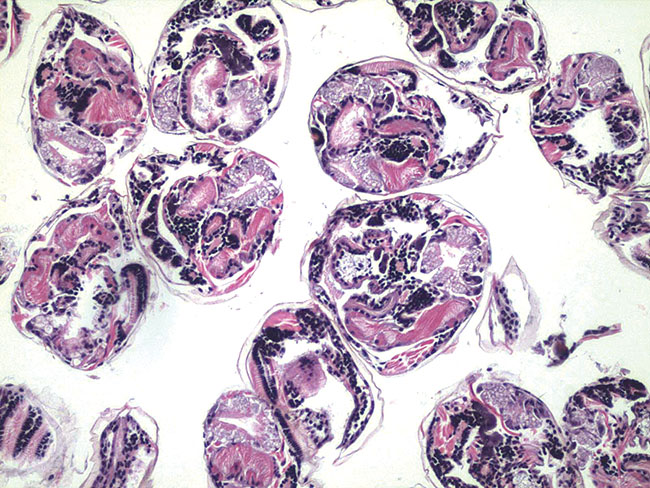 Under the microscope: Sections of juvenile hard clam tissues. The next phase of the research work, in participation with New York Sea Grant, will feature genomic sequencing at Stony Brook University. (Credit: Bassem Allam)
Under the microscope: Sections of juvenile hard clam tissues. The next phase of the research work, in participation with New York Sea Grant, will feature genomic sequencing at Stony Brook University. (Credit: Bassem Allam) A consortium of scientists is on the brink of completing a genomic map of the hard clam, Mercenaria mercenaria, in an ongoing offensive against QPX disease and the effects of extreme heat.
“It is 97 percent drafted,” says lead researcher Dr. Bassem Allam of Stony Brook University in New York.
Allam follows years of studies on how to reverse the shrinking clam populations along the Eastern Seaboard. He says this latest collaboration among five states and international stakeholders promises to finally yield “superior clam stocks.”
While oyster farming has been thriving, harvests of hard clams have faltered. According to the Aquaculture Census of the U.S. Department of Agriculture (USDA), the number of hard clams sold declined more than 10 percent between 2005 and 2018, while that of Eastern oysters shot up almost 60 percent.
“It’s really a contrast,” says Allam, attributing the decline to market constraints and natural mortality, like predation and disease.
Parasite unknown
Still, clams rank second among the most valuable species in U.S. aquaculture, according to the National Oceanic and Atmospheric Administration (NOAA). Hard clams in particular, which are also known as quahogs, are extensively cultured along the Atlantic coast. They are the most important marine resource in several states, including New York, Virginia and Florida.
But over the years, growers have suffered setbacks during disease outbreaks and heat waves.
For example, QPX (“quahog parasite unknown”) disease was first observed in Canada in 1959 and has since ravaged farms in Massachusetts, Virginia, New Jersey, New York, Nova Scotia and Prince Edward Island. Marked by chipped shell edges, the disease causes swelling and discoloration of the clam’s body wall and, in fact, can be fatal to the bivalve.
With such significant economic consequences, defeating QPX has been a high priority for aquaculture researchers.
The good news, Allam says, is that the threats of both a harsh environment and QPX disease can be offset by selective breeding.
Researching resistance
In a study funded by the USDA in 2016, Allam and his colleagues compared whether genetically distinct clam stocks resisted QPX disease. They were successful at identifying heritable traits associated with disease resistance in what Allam calls “a draft genome.”
They then selectively bred lines in New York during the summer of 2018 and deployed them in Massachusetts and New Jersey a year later to study the performance of these markers “at the same time, same place.”
The new research, funded by NOAA in 2019 under the National Aquaculture Initiative, will take advantage of single nucleotide polymorphism (SNPs).
“One single change in the DNA sequence is very important because it’s going to lead, ultimately, to different functional traits — to whether a hard clam is disease resistant or susceptible,” says Allam. “So, there is real hope down the line.”
The project, led by Allam in participation with New York Sea Grant, brings together scientists from five Atlantic coast states from Florida to Massachusetts, as well as contributions from Sorbonne Université in France. Clams for the study have been coming in from Florida to Prince Edward Island, and as far away as the U.K.
In the upcoming months, the team will complete the sequencing and assembly of the hard clam genome. From there, they will develop a genotyping platform or SNP array that can identify novel genetic markers associated with key traits.
The next step will be a validation period to audit resistance to QPX disease and heat stress.
These activities should take researchers well into 2022, but in the end, the platform will allow growers of hard clams to select traits relevant to various regions, including adaptability to salinity levels and tolerance of ocean acidification.
Print this page







